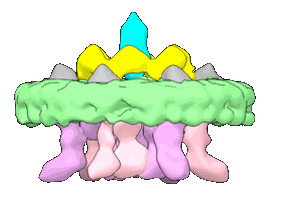
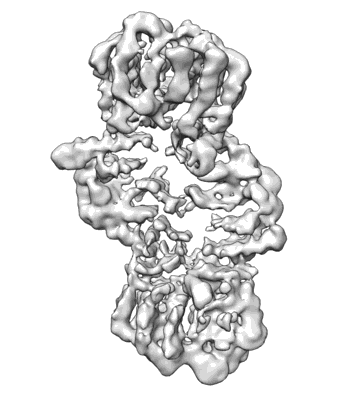
CryoEM structure of the TIR domain from AbTir in complex with 3AD
Resolution: 2.74 Å
EM Method: Helical reconstruction
Fitted PDBs: 7uxu
Q-score: 0.636
Manik MK, Shi Y, Li S, Zaydman MA, Damaraju N, Eastman S, Smith TG, Gu W, Masic V, Mosaiab T, Weagley JS, Hancock SJ, Vasquez E, Hartley-Tassell L, Kargios N, Maruta N, Lim BYJ, Burdett H, Landsberg MJ, Schembri MA, Prokes I, Song L, Grant M, DiAntonio A, Nanson JD, Guo M, Milbrandt J, Ve T, Kobe B
Science (2022) 377 pp. eadc8969-eadc8969 [ Pubmed: 36048923 DOI: doi:10.1126/science.adc8969 ]
- [[(2~{r},3~{s},4~{r},5~{r})-5-(6-aminopurin-9-yl)-3,4-bis(oxidanyl)oxolan-2-yl]methoxy-oxidanyl-phosphoryl] [(2~{r},3~{s},4~{r},5~{r})-5-(8-azanylisoquinolin-2-yl)-3,4-bis(oxidanyl)oxolan-2-yl]methyl hydrogen phosphate (686 Da, Ligand)
- Filament of the abtir tir domain in complex with 3ad (Complex from Acinetobacter baumannii)
- Molecular chaperone tir (15 kDa, Protein from Acinetobacter baumannii)
Porphyromonas gingivalis type IX secretion system
Song L, Perpich JD, Wu C, Doan T, Nowakowska Z, Potempa J, Christie PJ, Cascales E, Lamont RJ, Hu B
PNAS (2022) 119 pp. e2119907119-e2119907119 [ Pubmed: 35471908 DOI: doi:10.1073/pnas.2119907119 ]
- T9ss (Complex from Porphyromonas gingivalis)
Song L, Perpich JD, Wu C, Doan T, Nowakowska Z, Potempa J, Christie PJ, Cascales E, Lamont RJ, Hu B
PNAS (2022) 119 pp. e2119907119-e2119907119 [ Pubmed: 35471908 DOI: doi:10.1073/pnas.2119907119 ]
- T9ss (Complex from Porphyromonas gingivalis)
Song L, Perpich JD, Wu C, Doan T, Nowakowska Z, Potempa J, Christie PJ, Cascales E, Lamont RJ, Hu B
PNAS (2022) 119 pp. e2119907119-e2119907119 [ Pubmed: 35471908 DOI: doi:10.1073/pnas.2119907119 ]
- T9ss (Complex from Porphyromonas gingivalis)
pKM101-T4SS outer membrane core complex
Khara P, Song L, Christie PJ, Hu B
Mbio (2021) 12 pp. e0246521-e0246521 [ DOI: doi:10.1128/mBio.02465-21 Pubmed: 34634937 ]
pKM101-T4SS inner membrane complex
Khara P, Song L, Christie PJ, Hu B
Mbio (2021) 12 pp. e0246521-e0246521 [ DOI: doi:10.1128/mBio.02465-21 Pubmed: 34634937 ]
SARS-CoV-2 S-6P in complex with 9 Fabs
Resolution: 3.16 Å
EM Method: Single-particle
Fitted PDBs: 7e8c
Q-score: 0.39
Cao Y, Yisimayi A, Bai Y, Huang W, Li X, Zhang Z, Yuan T, An R, Wang J, Xiao T, Du S, Ma W, Song L, Li Y, Li X, Song W, Wu J, Liu S, Li X, Zhang Y, Su B, Guo X, Wei Y, Gao C, Zhang N, Zhang Y, Dou Y, Xu X, Shi R, Lu B, Jin R, Ma Y, Qin C, Wang Y, Feng Y, Xiao J, Xie XS
Cell Res (2021) 31 pp. 732-741 [ Pubmed: 34021265 DOI: doi:10.1038/s41422-021-00514-9 ]
- 604 h (23 kDa, Protein from Homo sapiens)
- Sars-cov-2 s (Complex from Homo sapiens)
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- N9 h (24 kDa, Protein from Homo sapiens)
- 604 l (23 kDa, Protein from Homo sapiens)
- 368-2 l (23 kDa, Protein from Homo sapiens)
- 368-2 h (24 kDa, Protein from Homo sapiens)
- Sars-cov-2 s-6p in complex with 9 fabs (Complex from Severe acute respiratory syndrome coronavirus 2)
- N9 l (22 kDa, Protein from Homo sapiens)
- 604 h, 604 l, n9 h, 368-2 l, 368-2 h, n9 l (Complex)
SARS-CoV-2 NTD in complex with N9 Fab
Resolution: 3.18 Å
EM Method: Single-particle
Fitted PDBs: 7e8f
Q-score: 0.592
Cao Y, Yisimayi A, Bai Y, Huang W, Li X, Zhang Z, Yuan T, An R, Wang J, Xiao T, Du S, Ma W, Song L, Li Y, Li X, Song W, Wu J, Liu S, Li X, Zhang Y, Su B, Guo X, Wei Y, Gao C, Zhang N, Zhang Y, Dou Y, Xu X, Shi R, Lu B, Jin R, Ma Y, Qin C, Wang Y, Feng Y, Xiao J, Xie XS
Cell Res (2021) 31 pp. 732-741 [ Pubmed: 34021265 DOI: doi:10.1038/s41422-021-00514-9 ]
- 604 h (23 kDa, Protein from Homo sapiens)
- 368-2 l (23 kDa, Protein from Homo sapiens)
- N9 h (24 kDa, Protein from Homo sapiens)
- 604 l (23 kDa, Protein from Homo sapiens)
- Sars-cov-2 ntd and rbd (Complex from Homo sapiens)
- Spike protein s1 (25 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Spike protein s1 (33 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- N9, 368-2 and 604h fab (Complex)
- 368-2 h (24 kDa, Protein from Homo sapiens)
- N9 l (22 kDa, Protein from Homo sapiens)
- Sars-cov-2 ntd in complex with n9 fab (Complex from SARS coronavirus Tor2)
Structural basis for control of antibiotic production by bacterial hormones
Zhou S, Bhukya H, Malet N, Harrison PJ, Rea D, Belousoff MJ, Venugopal H, Sydor PK, Styles KM, Song L, Cryle MJ, Alkhalaf LM, Fulop V, Challis GL, Corre C
Nature (2021) 590 pp. 463-467 [ DOI: doi:10.1038/s41586-021-03195-x Pubmed: 33536618 ]
- Protein-dna complex (114 kDa, Complex from Streptomyces coelicolor A3(2))
- Protein mmfr (Protein from Streptomyces coelicolor A3(2))
- Synthetic dna (DNA from Streptomyces coelicolor A3(2))
BD23-Fab in complex with the S ectodomain trimer
Resolution: 3.84 Å
EM Method: Single-particle
Fitted PDBs: 7byr
Q-score: 0.441
Cao Y, Su B, Guo X, Sun W, Deng Y, Bao L, Zhu Q, Zhang X, Zheng Y, Geng C, Chai X, He R, Li X, Lv Q, Zhu H, Deng W, Xu Y, Wang Y, Qiao L, Tan Y, Song L, Wang G, Du X, Gao N, Liu J, Xiao J, Su XD, Du Z, Feng Y, Qin C, Qin C, Jin R, Xie XS
Cell (2020) 182 pp. 73-84.e16 [ DOI: doi:10.1016/j.cell.2020.05.025 Pubmed: 32425270 ]
- Sars-cov-2 spike glycoprotein (Complex from SARS coronavirus 2)
- Ab23-fab (Complex from Homo sapiens)
- Spike glycoprotein (138 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Ab23-fab-light chain (11 kDa, Protein from Homo sapiens)
- Bd23-fab in complex with the s ectodomain trimer (Complex)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Ab23-fab-heavy chain (13 kDa, Protein from Homo sapiens)